MUSCLE or one of the Clustal algorithms like ClustalW. Most programs will align 3 or more sequences at a time and will require a different algorithm e.g. For comparing 2 sequences you’ll need to perform a “pairwise” alignment. You must have a minimum of 2 sequences to perform an alignment. We’re going to take a look at just the basics of sequence alignment to get you started. Whether you’re employing sequencing gels, Sanger-based methods, or the latest in pyrosequencing or ion torrent technologies, obtaining, manipulating and analyzing your sequences has never been easier. So although both the initial querry and the comparison database are nucleotide sequences, TBLAST-X results are provided as protein sequence alignments.Fortunately, those of us who have learned how to sequence know that aligning sequences is a lot easier and less time consuming than creating them. Each of the resulting six protein sequences is compared against the protein sequences resulting from the in silico 6 frame translation of a selected nucleotide sequence database. TBLAST-X (Nucleotide Querry → 6 Frame Translation vs. 6 Frame Translation of a Nucleotide Sequence DB BLAST)Ī user-provided nucleotide sequence is used to generated protein sequences from an in silico 6 frame translation. 6 Frame Translation of a Nucleotide Sequence DB BLAST)Ī user provided protein sequence is compared against the protein sequences resulting from the in silico 6 frame translation of a selected nucleotide sequence database. Each of the resulting six protein sequences is compared against a selected protein sequence database. Protein Sequence DB BLAST)Ī user-provided nucleotide sequence is used to generated protein sequences from an in silico 6 frame translation. Protein Sequence DB BLAST)Ī user-provided protein sequence is compared against a selected protein sequence database.īLAST-X (Nucleotide Querry → 6 Frame Translation vs. Nucleotide Sequence DB BLAST)Ī user-provided nucleotide sequence is compared against a selected nucleotide sequence database.īLAST-P (Protein Querry vs. 
The most commonly used alignment tools are provided with descriptions and links below:īLAST-N (Nucleotide Querry vs. The BLAST suite provides separate interfaces for many different querry sequence/database comparisons. assembled genome sequences, expressed sequence tags (ESTs), NCBI genomes, patented protein sequences, protein database (pdb) proteins, etc.). The Basic Local Alignment Search Tool (BLAST) allows you to perform local alignments between a user-provided DNA, RNA or protein sequence and the sequences containined in a large number of curated sequence databases (e.g. Molecular systems biology 7(1):1-6.Ībout the Basic Local Alignment Search Tool (BLAST): Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Sievers F, Söding J, Thompson JD, Higgins DG, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R et al. You can download the appropriate application installer from here: The Clustal Omega program can also be downloaded for stand alone use on personal computers running Mac, Windows or Linux operating systems from the following location:ĭownloading the JalView Sequence Alignment Results Viewer:Īlthough Clustal Omega provides a serviceable text-based display of alignment results, the JalView alignment results viewer provides a much nicer way to do it. The application can be used online via hosted web servers at the following locations:Ĭlustal Omega (Web Server at Pasteur Mobyle Portal)ĭownloading the Application to Your Personal Computer: Uploaded sequence files are limited to a maximum of 4 MB. The current version of the software accepts a maximum of 4000 sequences. Clustal Omega is an improved version of that tool.Īccepted sequence formats are GCG, FASTA, EMBL, GenBank, PIR, NBRF, PHYLIP or UniProtKB/Swiss-Prot. 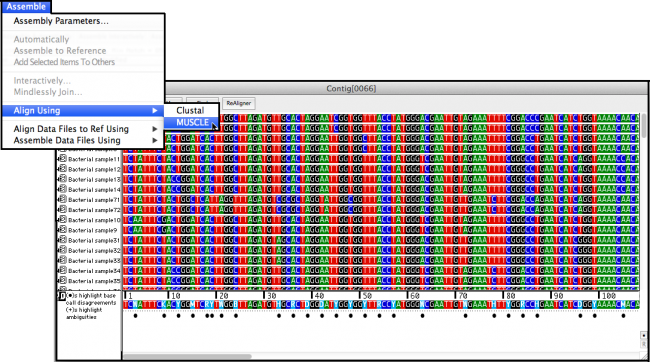
For many years, the previous version of the tool, Clustal W, was widely used for this kind of multiple sequence alignment. Clustal Omega is a multiple sequence alignment tool best used for aligning similar sequence regions between three or more RNA, DNA or protein sequences. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
